LentiFlash is a fast and safe gene editing delivery technology for Cre-loxP models

The use of the Cre-loxP system increases as technological developments broaden its applicability
The discovery and initial characterization of the Cre-loxP system came out from the works carried out by the scientist Nat L. Sternberg [1] .These insights and efforts made Cre-loxP an invaluable genetic tool. Dr. Brian Sauer developed Cre-loxP recombination that operated in both mitotic and non-mitotic cells, and was initially used in activating gene expression in mammalian cell lines [2].
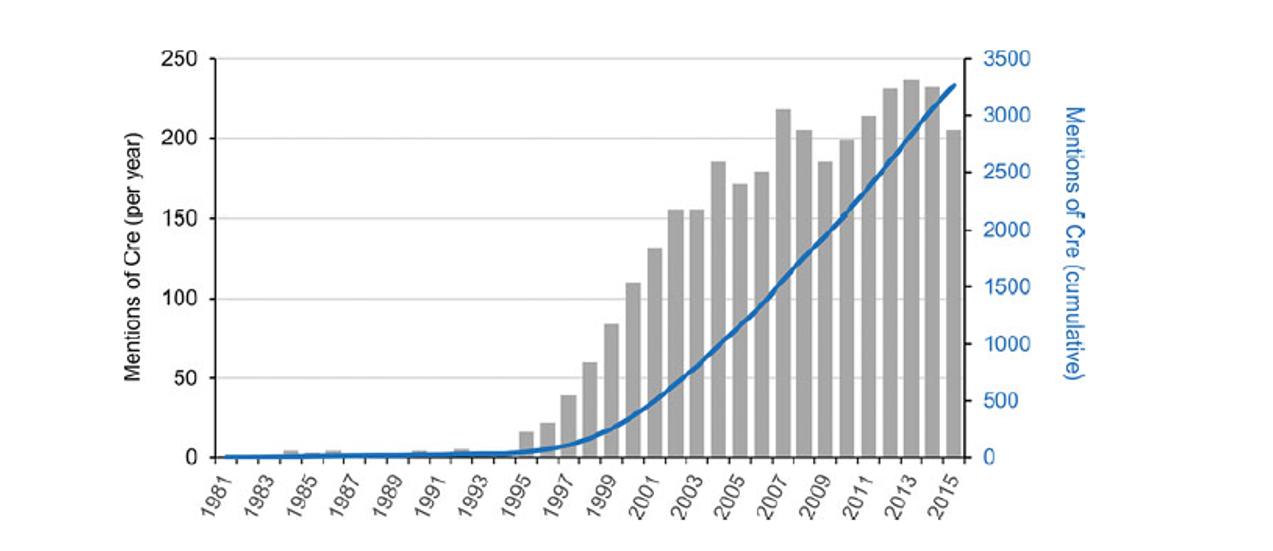
Finally, Dr. Jamey Marth demonstrated that Cre-loxP recombination could be used to delete loxP-flanked chromosomal DNA sequences at high efficiency in specific developing T-cells of transgenic animals [3]. Since the development of the Cre-loxP system as a research tool, more than 3,000 reports have been published utilizing the technology [4] (Figure 1).
Figure 1. The use of the Cre-loxP system increases as technological developments broaden its applicability. Grey bar graph (left y-axis) and blue line graph (right y-axis) display yearly and cumulative mentions, respectively, of “Cre-loxP” in published scientific reports since its development (1981 to 2015). The database used was Pubmed, and the search terms were “loxp” and “cre”.
Cre-loxP recombination became ideal for performing genetic manipulation in vivo
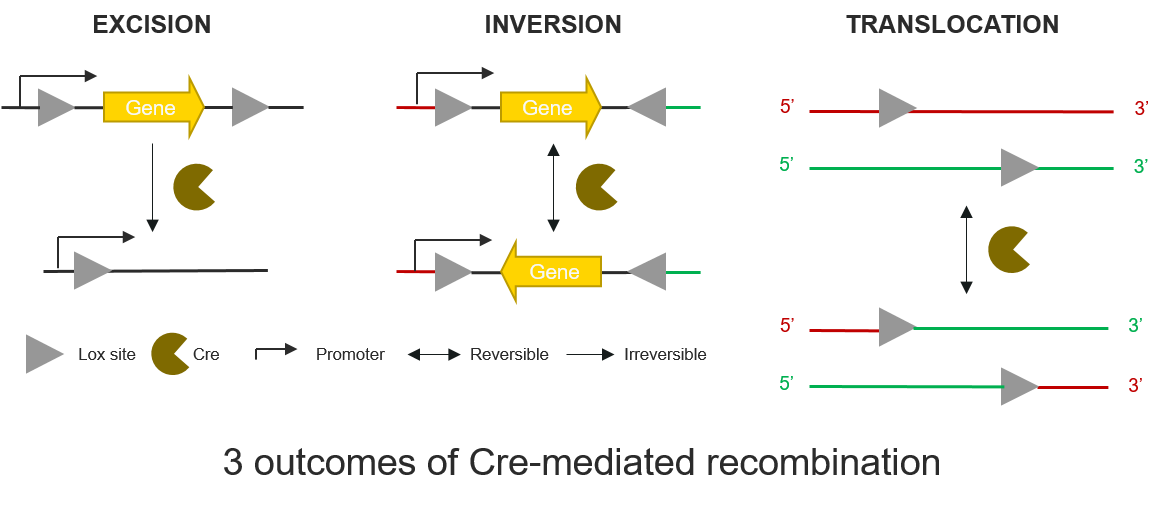
Cre-loxP system is a powerful genetic tool which can be used to control gene expression by excision, inversion or translocation to the genome [4]. According to the orientation of the lox sites, there are 3 types of possible recombination at the lox-flanked regions of DNA. These lox regions are often said to be floxed:
- if the lox sites are in the same direction, the floxed region will be excised.
- if the lox sites are in the opposite direction, the floxed region will be reversed.
- if the lox sites are on 2 different DNA regions, the recombination will produce a translocation event.
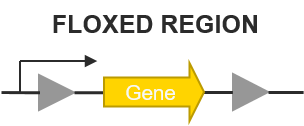
Figure 2. Common representation of a cassette containing a floxed region.
Figure 3. Schematic representation of the 3 possible types of recombination of a floxed region, according to the Lox sites orientation.
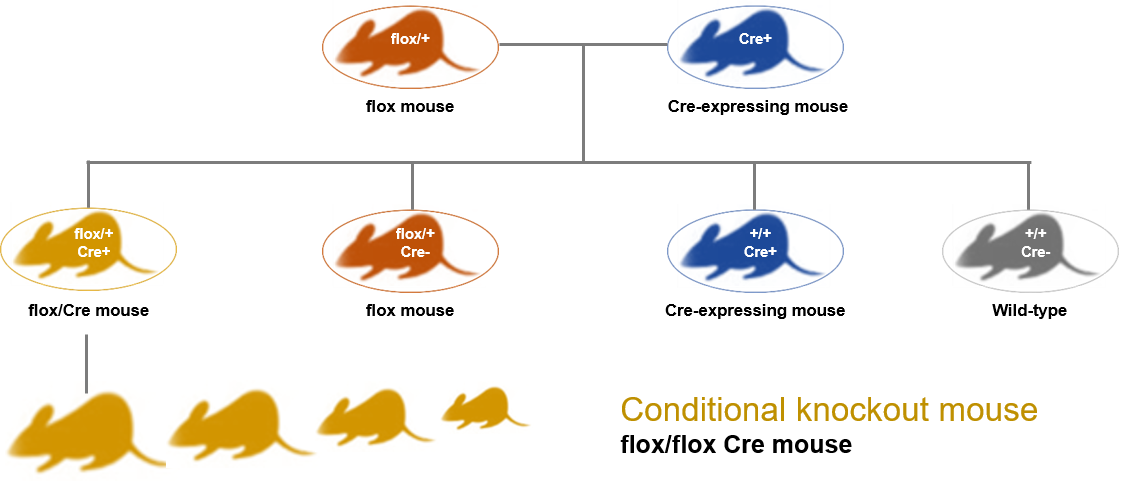
These predictable recombinations are used to generate conditional KO and fusion reporter models for studying gene function and regulation. However, this system of genetically engineered mouse models remains technically complex. LoxP DNA sites must be first introduced to flank the gene of interest into the genome of animal (Floxed region), while transgenic Cre-driver mice are also generated. Then, the selection and production of a homozygous transgenic generation is required, allowing conditional knockouts via the expression of Cre recombinase.
Figure 4. Schematic representation of the generation of genetically engineered mouse models using the Cre-loxP system.
LentiFlash addresses the major obstacles of Cre-delivery system
In vivo, conditional mutagenesis requires developing animal breeding strategies, which can become time-consuming and expensive. The level of heterogeneity of these crossings due to the choice of mouse strain and also to the nature and the function of the targeted genes can also be laborious. Nevertheless, editing using Cre addition is predictable and robust to remove or reverse floxed sequences.
In fact, the major obstacle is the lack of robust and easy-to-use Cre-delivery systems, ensuring the rapid and efficient delivery of Cre recombinase into loxP mice models. To address this issue, Flash Therapeutics combined the efficiency of the LentiFlash® RNA delivery tool to the Cre-loxP gene editing technology.
The data exposed below demonstrate that LentiFlash® allows the efficient delivery of the Cre recombinase to generate 2 types of recombination, excision and inversion, therefore usable in many genetically engineered Cre-loxP mouse models.
LentiFlash delivers functional Cre mRNA locally in vivo in many tissues
Thanks to LentiFlash® the CRE system can be delivered as efficiently as the classical lentiviral vectors but in a non-integrative manner. The following experiments show how the Flash Therapeutics teams obtained good delivery results of Cre mRNA in different tissues:
Cre-loxP recombination generating excision
Experiment 1: LentiFlash® Cre injected locally in muscle of ROSA26-YFP reporter mice
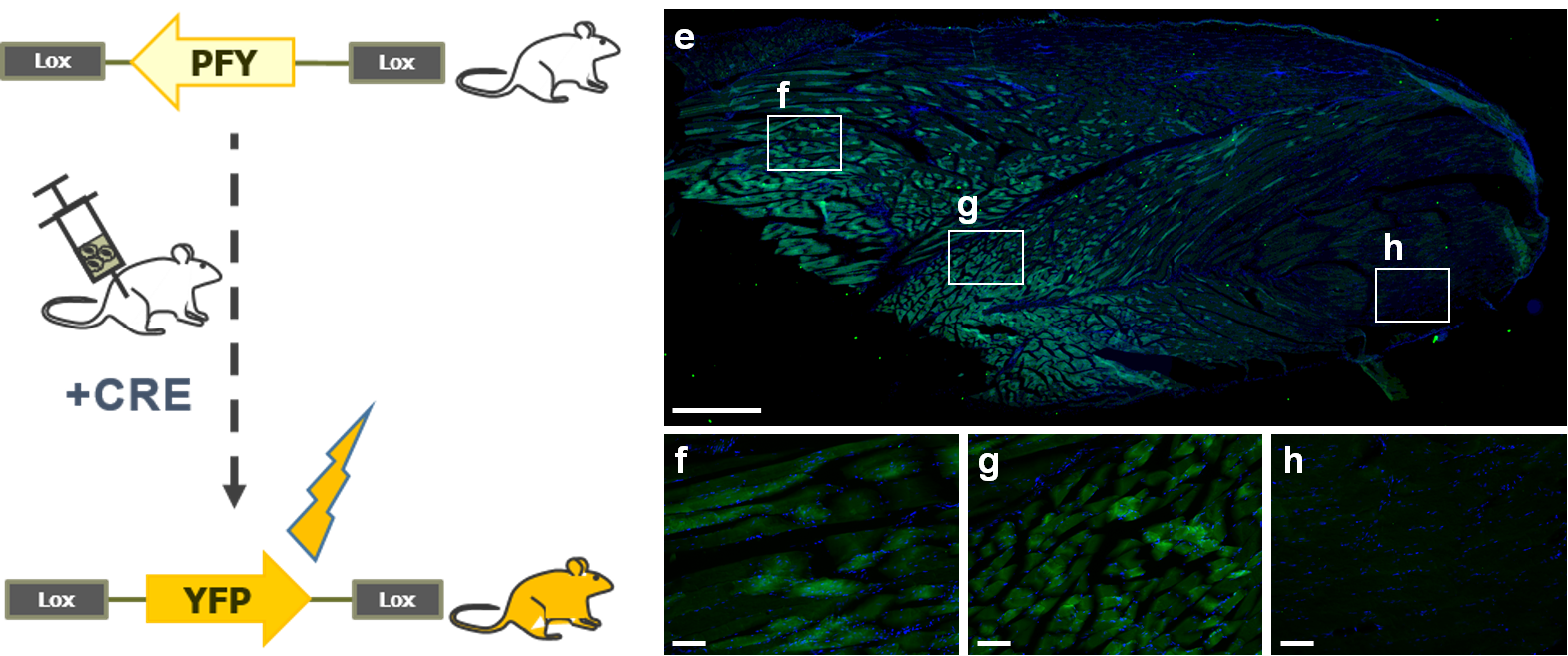
In this experiment, published in 2015 by (Prel et al. Mol.Ther.) LentiFlash® Cre (1E+07 physical particles in 5µl) was locally injected into the biceps femoris muscle of five ROSA26-YFP reporter mice. Injected samples were collected and analyzed 7 days post-injection. As shown in Figure 5, this data highlights the efficiency of the non-integrative LentiFlash® Cre system, which conferred expression of the YFP in a wide zone of injected muscle. This in vivo data confirmed the potency of LentiFlash® to deliver functional Cre mRNA locally.
Figure 5. In vivo analysis for YFP expression in ROSA 26 YFP mice after injection of LentiFlash® Cre. YFP immuno-staining (green) on 20 μm thick cryosections of injected muscles. Nuclei were stained with Hoechst 33342 (blue). Mice were injected with LentiFlash® Cre (E–G); internal negative control is shown (H). Phosphate-buffered saline–injected muscle were not stained (data not shown). Scale bars: E = 1 mm, F–H = 100 μm.
Experiment 2: LentiFlash® Cre injected locally in the trachea of mTmG reporter mice
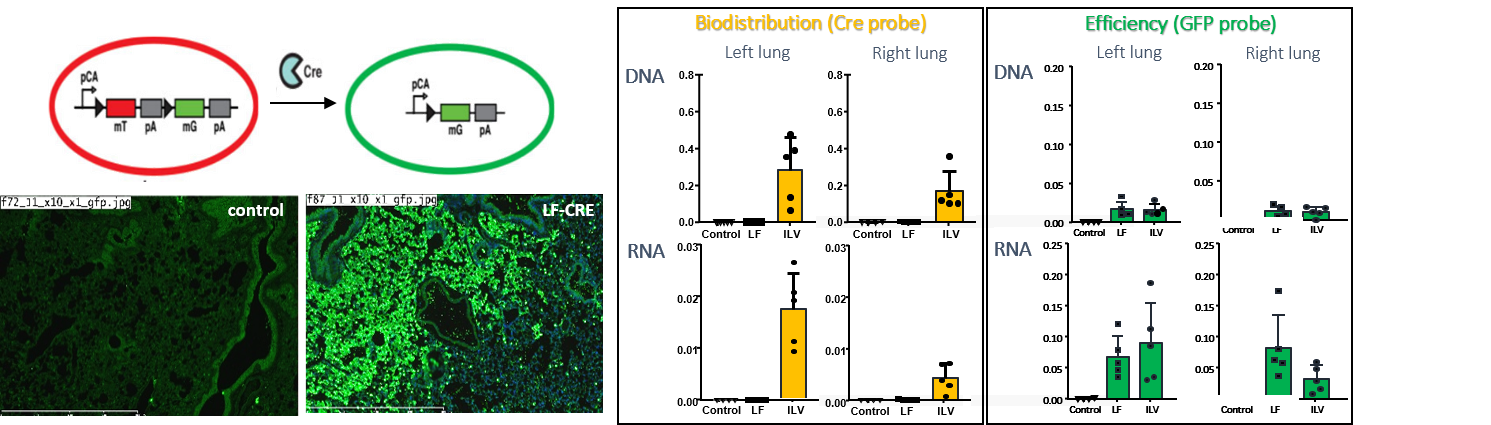
In a collaboration with the “Institut Clinique de la Souris” (ICS) in Strasbourg (France), 25µl of LentiFlash® Cre was injected locally by intra tracheal route into mTmG reporter mice and analyzed 2 weeks after injection. As shown in figure 6a, LentiFlash® Cre provides high deletion efficiency of the floxed-tdTomato region as shown by the bright local expression of GFP reporter into the lungs. No excision is observed in other organs (data not shown).
Figure 6b reports ddPCR molecular analyses on RNA and DNA lung samples from mTmG reporter mice injected 2 weeks earlier either with LentiFlash® Cre or with integrative lentiviral vector, iLV-EF1-Cre (control). This data shows that :
- The Cre recombinase is detected only after ILV injections, as expected, as the LentiFlash particles lead to only a transient protein expression.
- GFP reporter is detected as efficiently after ILV or LentiFlash injections.
This data validates that a high deletion efficiency is obtained with the LentiFlash® with no residual expression of the Cre recombinase.
Figure 6. (a) In vivo analysis for GFP expression in mTmG reporter mice (n=5) 2 weeks after injection of LentiFlash® Cre. Lungs histology was made with GFP immuno-staining (green). (b) ddPCR analyses were performed on gDNA samples to detect gene integration and on RNA samples to quantify the gene expression in mTmG reporter mice (n=5) 2 weeks after injection either with a LentiFlash® Cre or an integrative lentiviral vector control, iLV-EF1-Cre.
Cre-loxP recombination generating inversion
Experiment 3: LentiFlash® Cre injected locally in muscle of GOI-GFP reporter mice for the generation of a vaccination model
In a collaboration with the National Veterinary School of Toulouse (ENVT) (France), we developed a model dedicated to the evaluation of a vaccine candidate. The expression of the protein targeted by the vaccine is induced locally in the genetically modified mice using LentiFlash® expressing the Cre recombinase. This murine model is thus designed to easily evaluate the efficacy of a therapeutic vaccine, in a controlled manner in time and space.
Our data shows that:
- Gene-s included within the Floxed cassette are efficiently inversed and thus expressed thanks to LentiFlash®-Cre injections
- LentiFlash®-Cre allows an efficient local expression of any gene of interest into any Floxed model.
This data validates that a high inversion efficiency is obtained with the LentiFlash®.
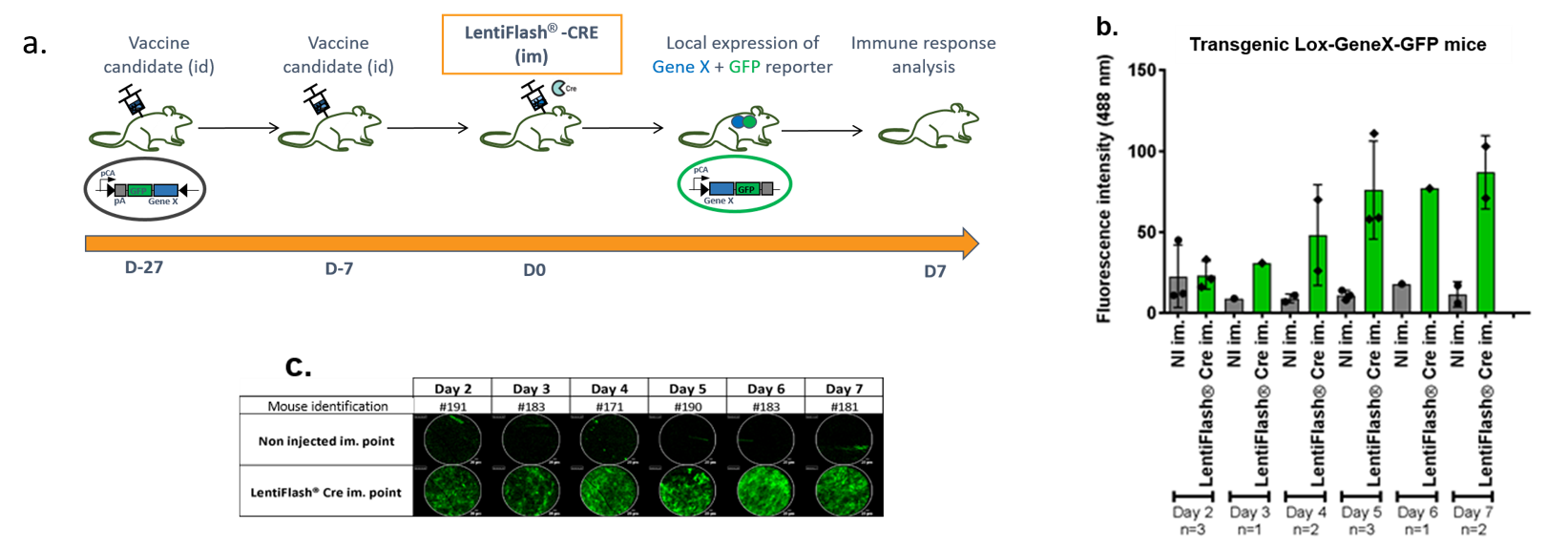
Figure 7. (a) Schematic representation of the strategy. Transgenic mice are floxed for an inverted cassette containing the GeneX fused to the GFP reporter protein. After vaccine priming and boost, transgenic mice are injected by intramuscular (im) route with the LentiFlash®-Cre. The detection of the local expression of GFP allows the follow-up of the protein X expression. It is performed with the Cellvizio® imaging system (MaunaKea technologies, France) from day 2 to day 7. Vaccine response is analyzed at day 7. (b) Analysis of the GFP fluorescence intensity (mean ± SD on the totality of each point all along the video, every day for every point. NI : Non Injected (c) Images of the local expression of GFP in the muscle.
To learn more about LentiFlash® Technology do not hesitate to visit our website
References:
[1] Yarmolinsky M. and Hoess R. (2015). The Legacy of Nat Sternberg: The Genesis of Cre-loxP Technology. Annu. Rev. Virol. 2:25–40
[2] Sauer B. (1987). Functional expression of the Cre-lox site-specific recombination system in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 7(6):2087‐2096
[3] Orban PC, Chui D, Marth JD. (1992). Tissue- and site-specific DNA recombination in transgenic mice. Proc Natl Acad Sci U S A. 89(15):6861‐6865
[4] McLellan, M.A., Rosenthal, N.A., and Pinto, A.R. (2017) Cre-loxP-mediated recombination: General principles and experimental considerations. Curr. Protoc. Mouse Biol. (2017) 7:1-12
[5] Channabasavaiah B.G.and Kent Lloyd K.C. (2019) Generating mouse models for biomedical research: technological advances. Disease Models and Mechanisms. 12:1-10